This past week we’ve been blogging about BigML’s new Principal Component Analysis (PCA) feature. In this post, we will continue on that topic and discuss some of the mathematical fundamentals of PCA, and reveal some of the technical details of BigML’s implementation.
A Geometric Interpretation
Let’s revisit our old friend the iris dataset. To simplify visualizations, we’ll just look at two of its variables: petal length and sepal length. Imagine now that we take some line in this 2-dimensional space and we sum the perpendicular distances between this line and each point in the dataset. As we rotate this line around the center of the dataset, the value of this sum changes. You can see this process in the following animation.
What we see here is that the value of this sum reaches a minimum when the line is aligned with direction in which the dataset is most “stretched out”. We call a vector that points in this direction the first “principal component” of the dataset. It is plotted on the left side of the figure below so you don’t need to make yourself dizzy trying to see it in the animated plot.
The number of principal components is equal to the dimensionality of the dataset, so how do we find the second one for our 2D iris data? We need to first cancel out the the influence of the first one. From each data point, we will subtract its projection along the first principal component. As this is the final principal component, we will be left with all the points in a neat line, so we don’t need to go spinning another vector around. This is the result shown on the right side of the figure below.
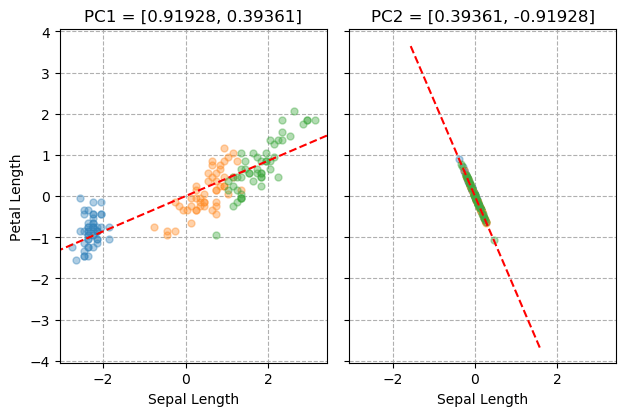
The importance of a principal component is usually measured by its “explained variance”. Our example dataset has a total variance of 3.80197. After we subtract the first principal component, the total variance is reduced to 0.14007. The explained variance for PC1 is therefore:
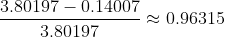
This process can be generalized for data with higher dimensions. For instance, starting with a 3D point cloud, subtracting the first principal component gives you all the points in a plane, and then the process for the finding the second and third components is the same as what we just discussed. However, for even a moderate number of dimensions, the process becomes extremely difficult to visualize, so we need to turn to some linear algebra tools.
Covariance Decomposition
Given a collection of data points, we can compute its covariance matrix. This is a quantification of how each variable in the dataset varies both individually and together with other variables. Our 2D iris dataset has the following covariance matrix:
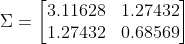
Here, sepal length and petal length are the first and second variables respectively. The numbers on the main diagonal are their variance. The variance of sepal length is several times that of petal length, which we can also see when we plot the data points. The value on the off diagonal is the covariance between the two variables. A positive value here indicates that they are positively correlated.
Once we have the covariance matrix, we can perform eigendecomposition on it to obtain a collection of eigenvectors and eigenvalues.
![\mathbf{e}_1 = [ 0.91928 , 0.39361] \qquad \lambda_1 = 3.66189 pca_eq3](https://blog.bigml.com/wp-content/uploads/2018/12/pca_eq3.gif?w=810)
![\mathbf{e}_2 = [0.39361, -0.91928] \qquad \lambda_2 = 0.14007 pca_eq4](https://blog.bigml.com/wp-content/uploads/2018/12/pca_eq4.gif?w=810)
We can now make a couple observations. First, the eigenvectors are essentially identical to the principal component directions we found using our repeated line fits. Second, the eigenvalues are equal to the amount of explained variance for the corresponding component. Since covariance matrices can be constructed for any number of dimensions, we now have a method that can be applied whatever size problem we wish to analyze.
Missing Values
When we move beyond the realm of toy datasets like iris, one of the challenges we encounter is how to deal with missing values, as the calculation of the covariance matrix does not admit those. One strategy could be to simply drop all the rows in your dataset that contain missing data. With a high proportion of missing values however, this may lead to an unacceptable loss of data. Instead, BigML’s implementation employs the unbiased estimator described here. First, we form a naive estimate by replacing all missing values with 0 and computing the covariance matrix as normal. Call this estimate 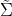 , and let
, and let ![]() be the same matrix with the off-diagonal elements set to 0. We also need to compute a parameter
be the same matrix with the off-diagonal elements set to 0. We also need to compute a parameter 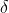 , which is the proportion of values which are not missing. This is easily derived from the dataset’s field summaries. The unbiased estimate is then calculated as:
, which is the proportion of values which are not missing. This is easily derived from the dataset’s field summaries. The unbiased estimate is then calculated as:
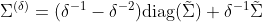
With our example data, we randomly erase 100 values. Since we have 150 2-dimensional datapoints, that gives us 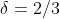 . The naive and corrected missing value covariance matrix estimates are:
. The naive and corrected missing value covariance matrix estimates are:
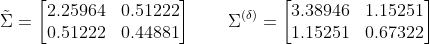
The corrected covariance matrix is clearly much nearer to “true” values.
Non-Numeric Variables
PCA is typically applied to numeric-only data, but BigML’s implementation incorporates some additional techniques which allow for the analysis of datasets containing other data types. With text and items fields, we explode them into multiple numeric fields, one per item in their tag cloud, where the value is the term count.
Categorical fields are slightly more involved. BigML uses a pair of methods called Multiple Correspondence Analysis (MCA) and Factorial Analysis of Mixed Data (FAMD). Each field is expanded into multiple 0/1 valued fields 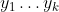 , one per categorical value. If p is the mean of each such field, then we compute the shifted and scaled value, using one of the two expressions below:
, one per categorical value. If p is the mean of each such field, then we compute the shifted and scaled value, using one of the two expressions below:
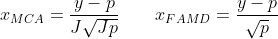
If the dataset consists entirely of categorical fields, the first equation is used, and J is the total number of fields. Otherwise, we have a mixed datatype dataset and we use the second equation. After these transformations are applied, we then feed the data into our usual PCA solver.
We hope this post has shed some light on the technical details of Principal Component Analysis, as well as highlight the unique technical features BigML brings to the table. To put your new-found understanding to work, head over to the BigML Dashboard and start running some analyses!
Want to know more about PCA?
For further questions and reading, please remember to visit the release page and join the release webinar on Thursday, December 20, 2018. Attendance is FREE of charge, but space is limited so register soon!
